TL;DR —
Supplementary figures showcase key findings from CheSS searches, including Tanimoto coefficients, feature cosine similarities, and token vector comparisons. These visuals highlight how different SMILES canonicalizations impact molecular search behavior, emphasizing structural vs. functional similarity in chemical discovery.
Authors:
(1) Clayton W. Kosonocky, Department of Molecular Biosciences, The University of Texas at Austin (clayton.kosonocky@utexas.edu);
(2) Aaron L. Feller, Department of Molecular Biosciences, The University of Texas at Austin (aaron.feller@utexas.edu);
(3) Claus O. Wilke, Department of Integrative Biology, The University of Texas at Austin and Corresponding Author (wilke@austin.utexas.edu);
(4) Andrew D. Ellington, Department of Molecular Biosciences, The University of Texas at Austin (ellingtonlab@gmail.com).
Table of Links
- Abstract & Introduction
- Methods
- Results and Discussion
- Determining Whether Canonicalization Impacts Search Behavior
- Explanation of Search Behavior & Drawbacks, Future Improvements, and Potential for Misuse
- Conclusion, Acknowledgements, Author Contributions, & more.
- Supplementary Figures
Supplementary Figures

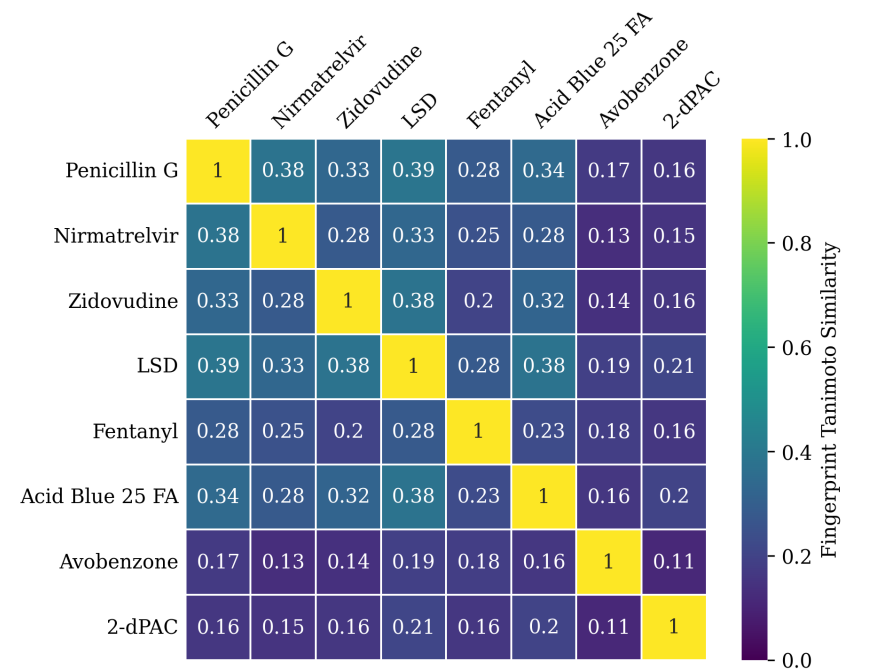
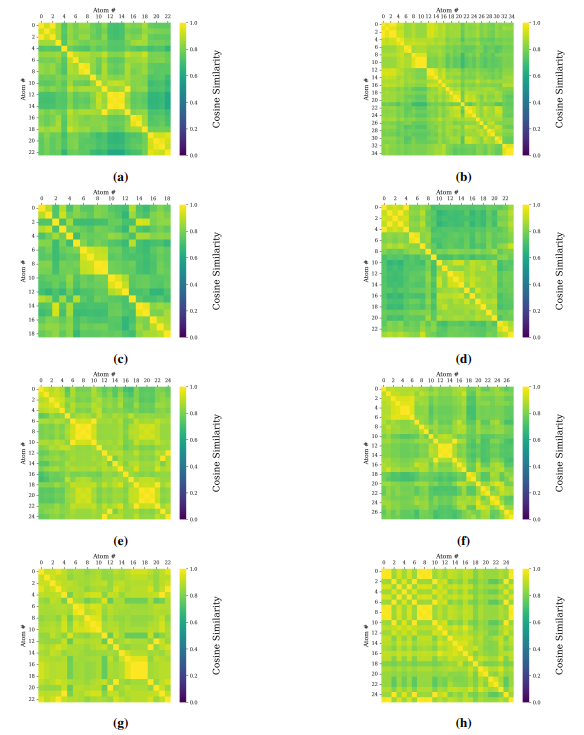
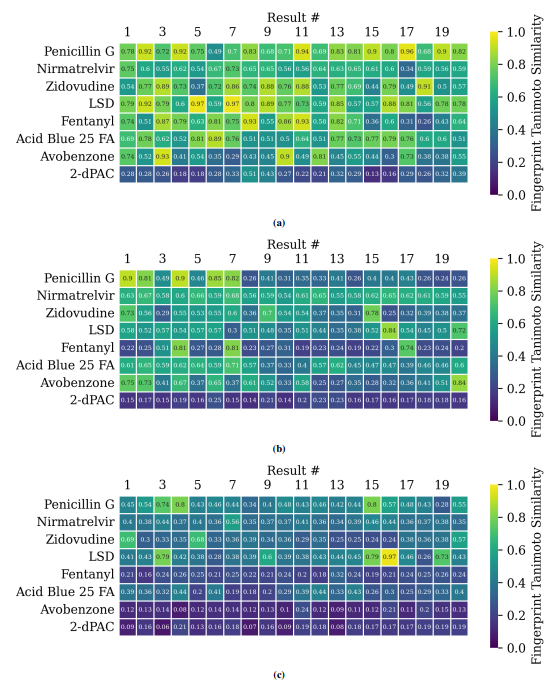
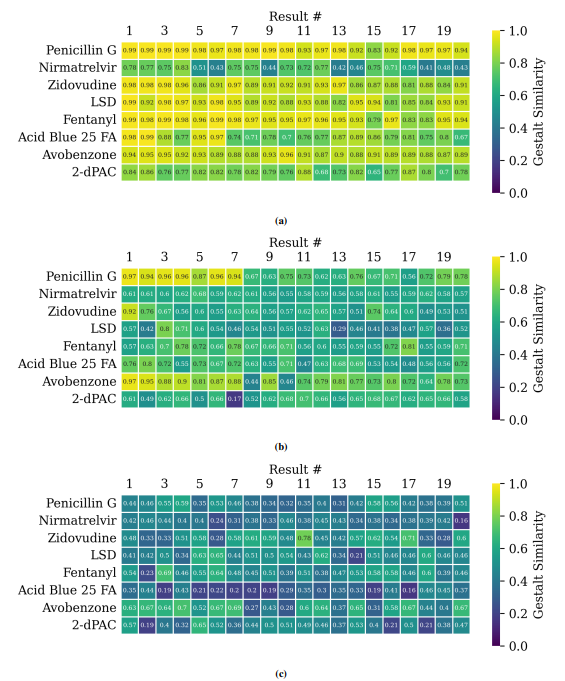
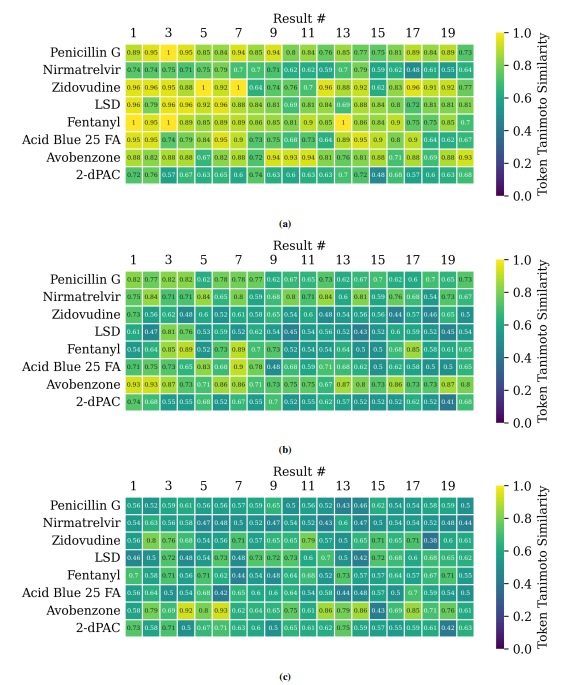
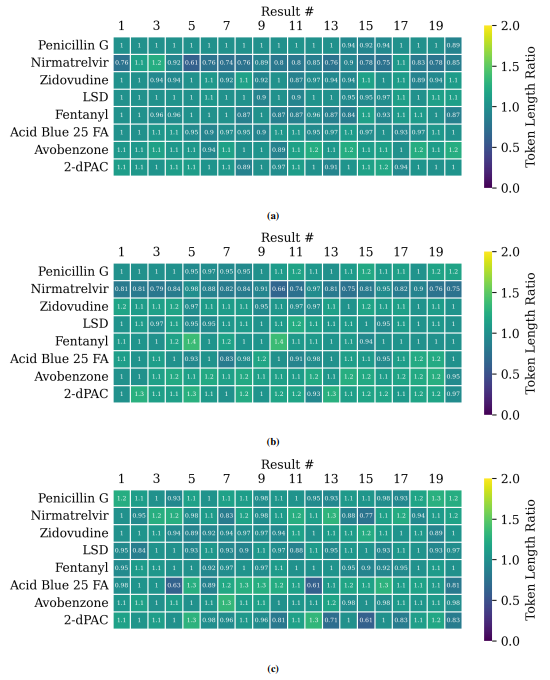
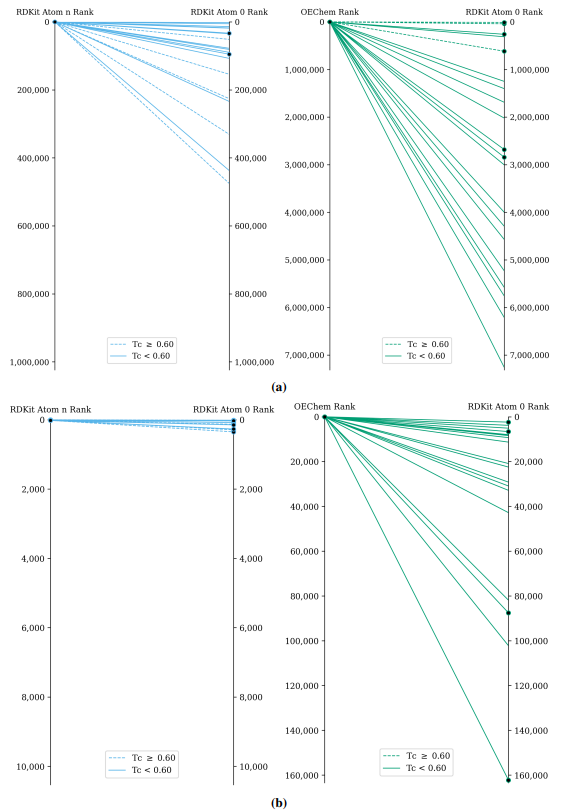
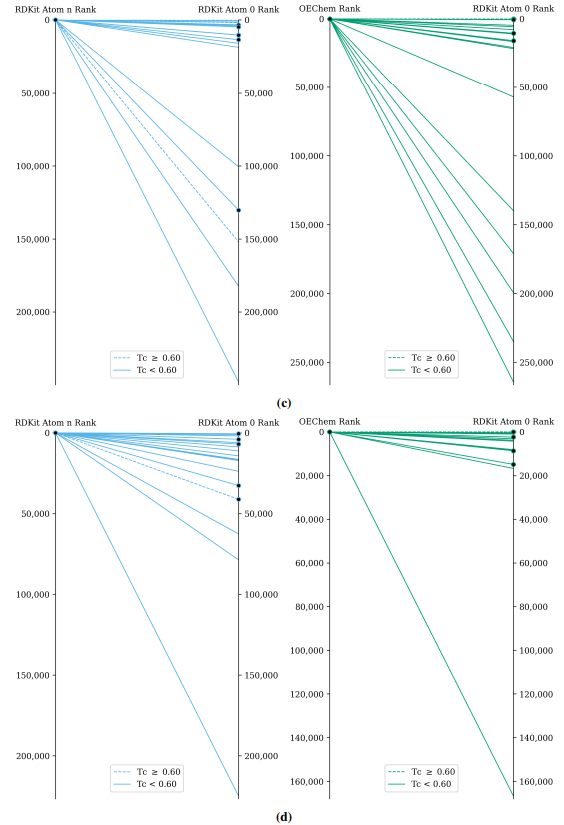
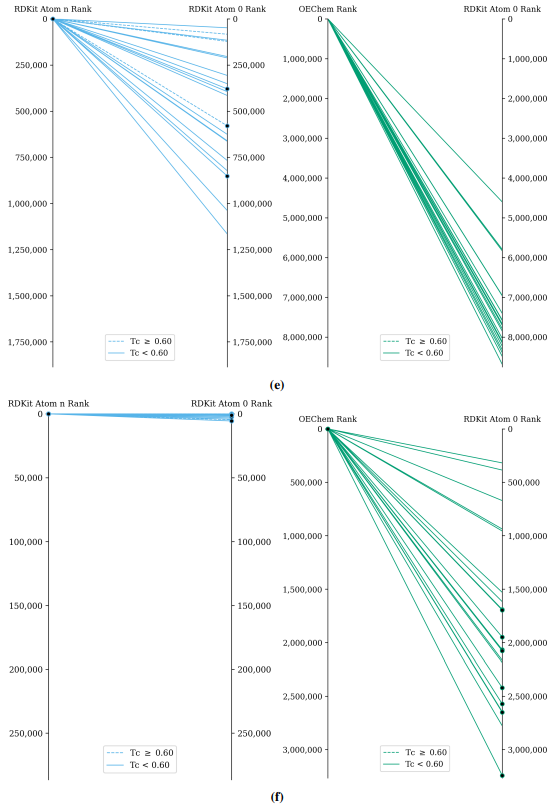
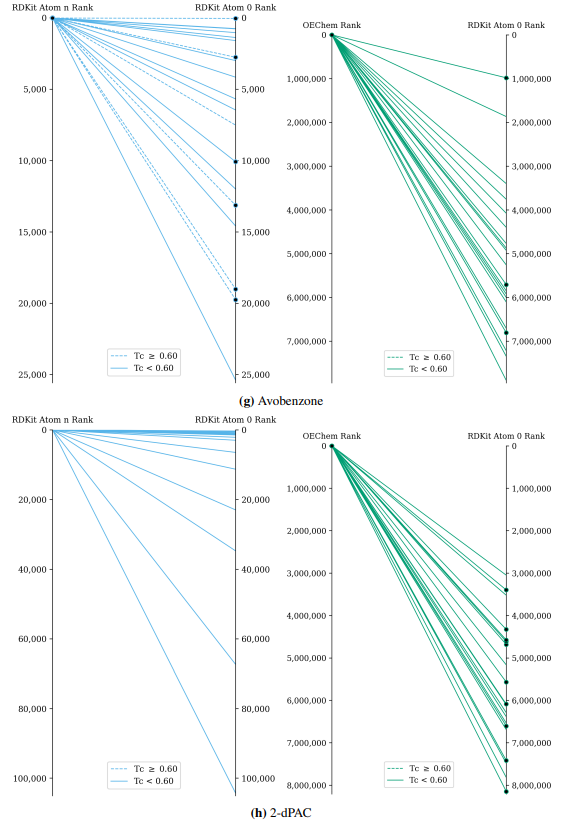
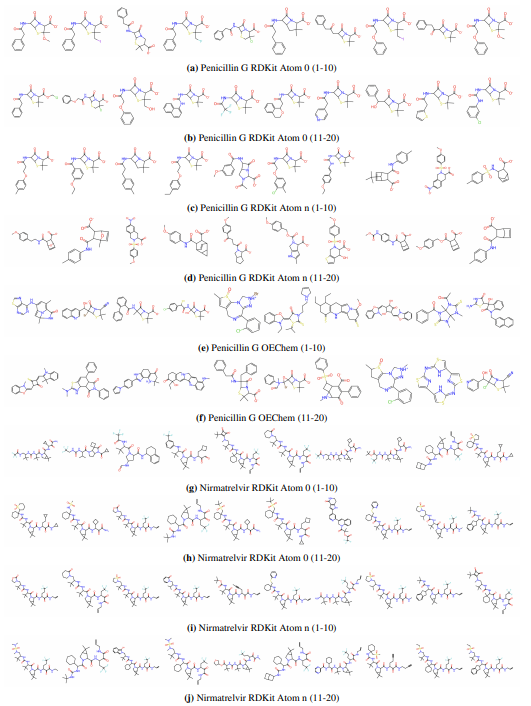
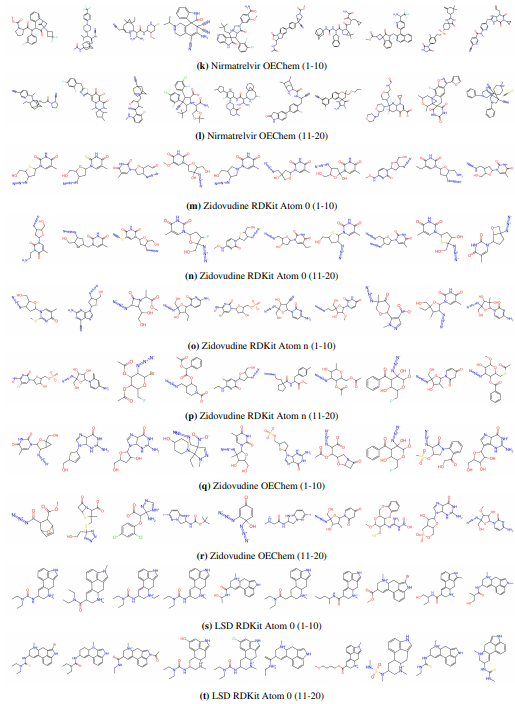
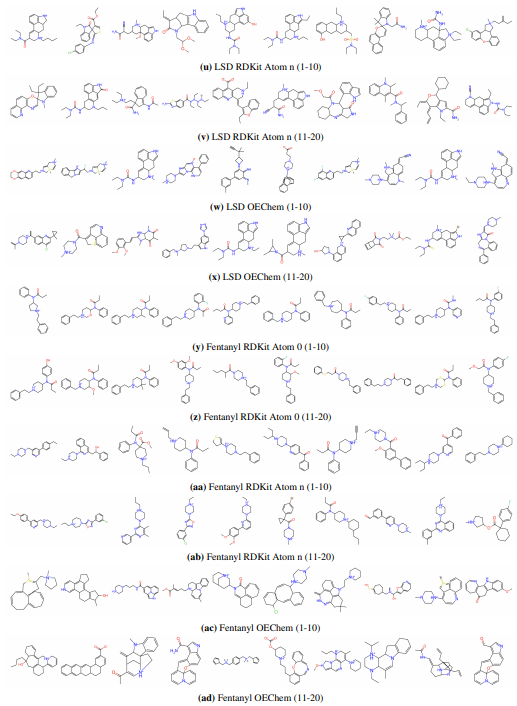
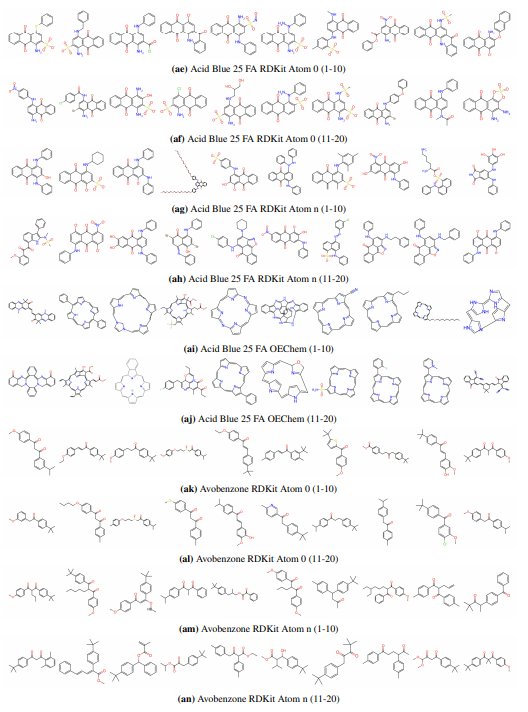
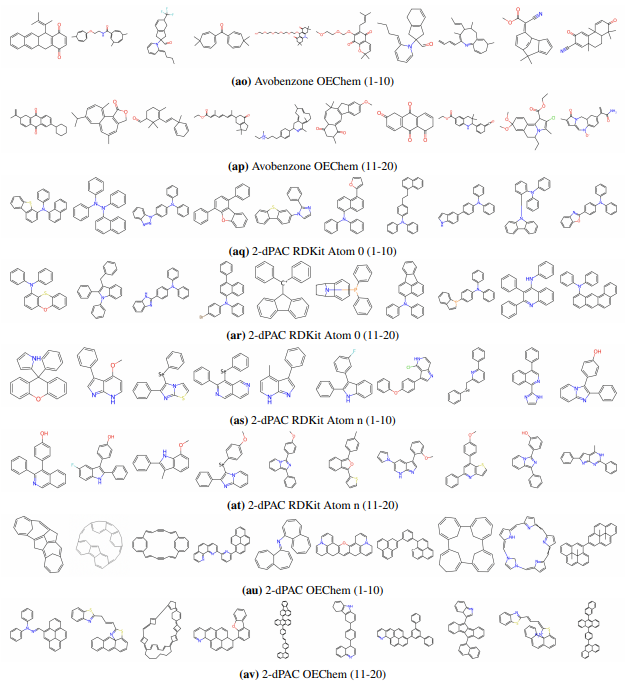
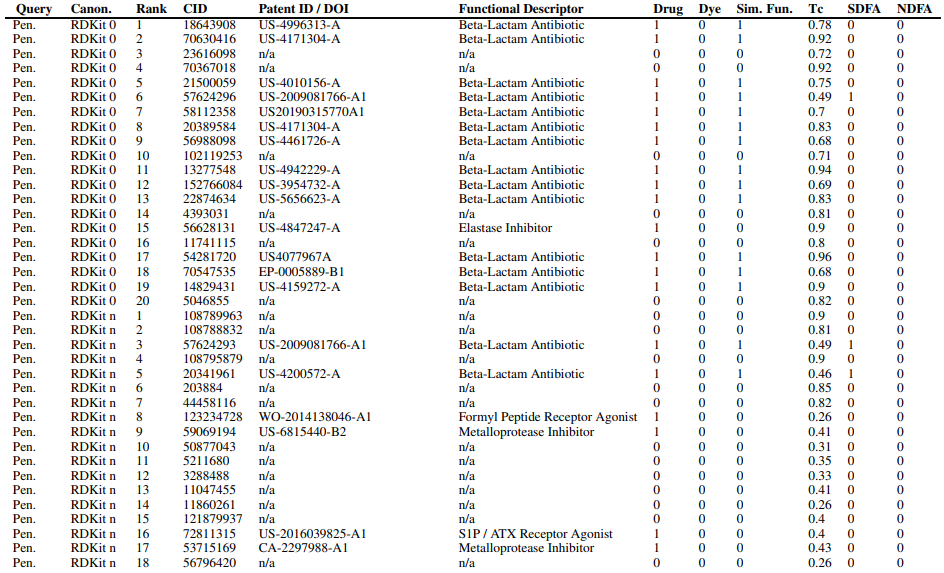
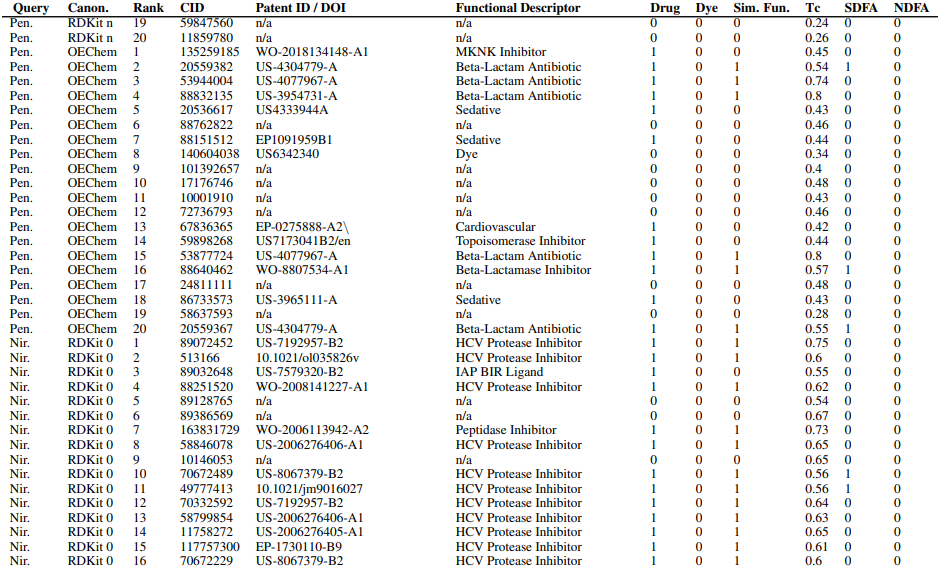
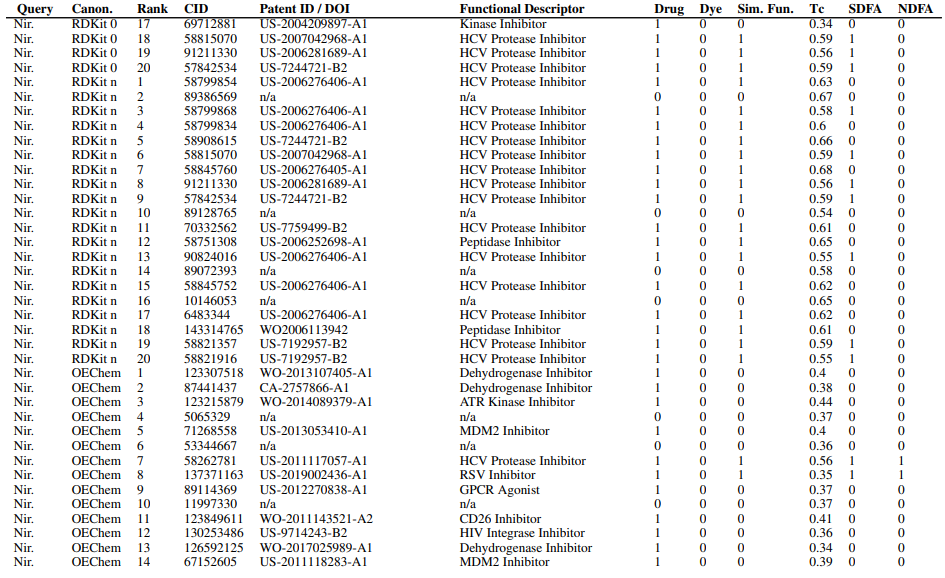
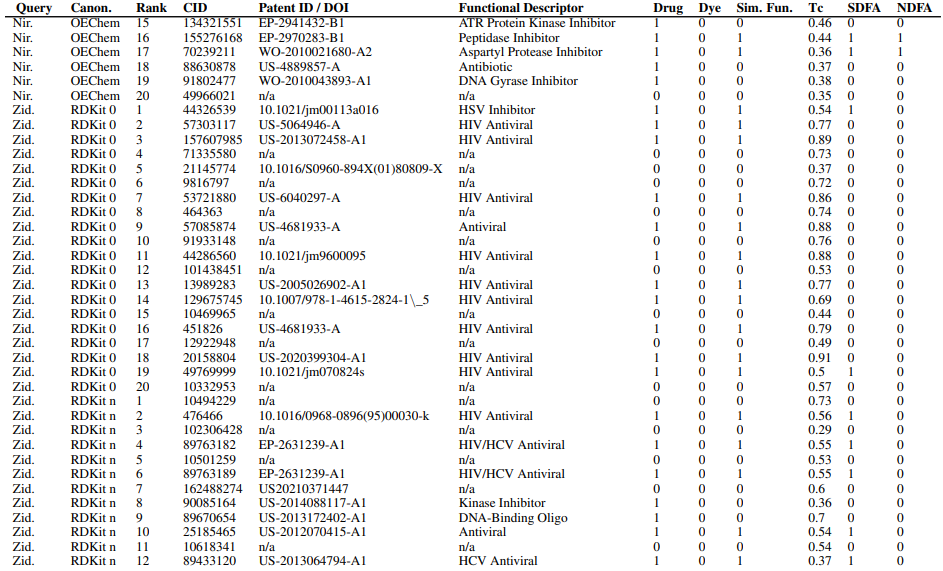
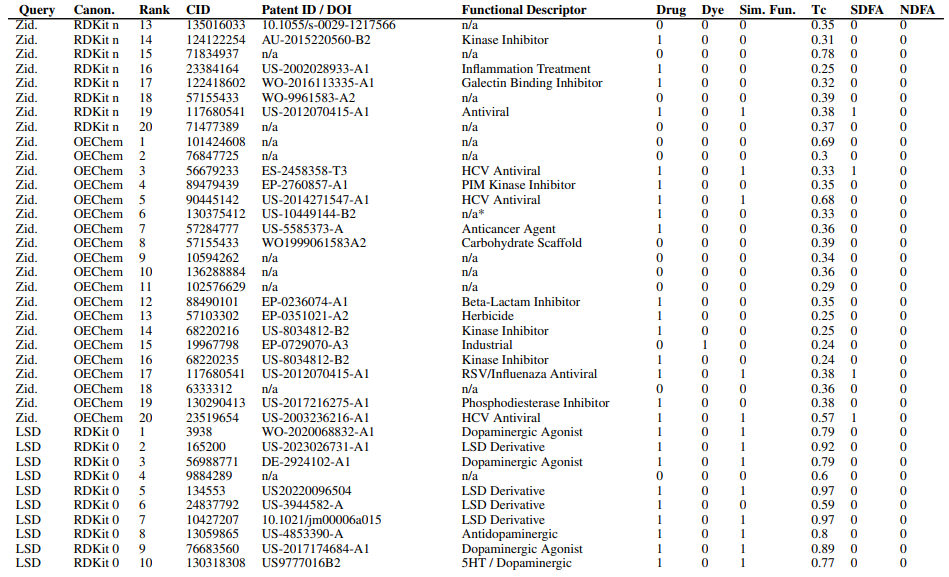
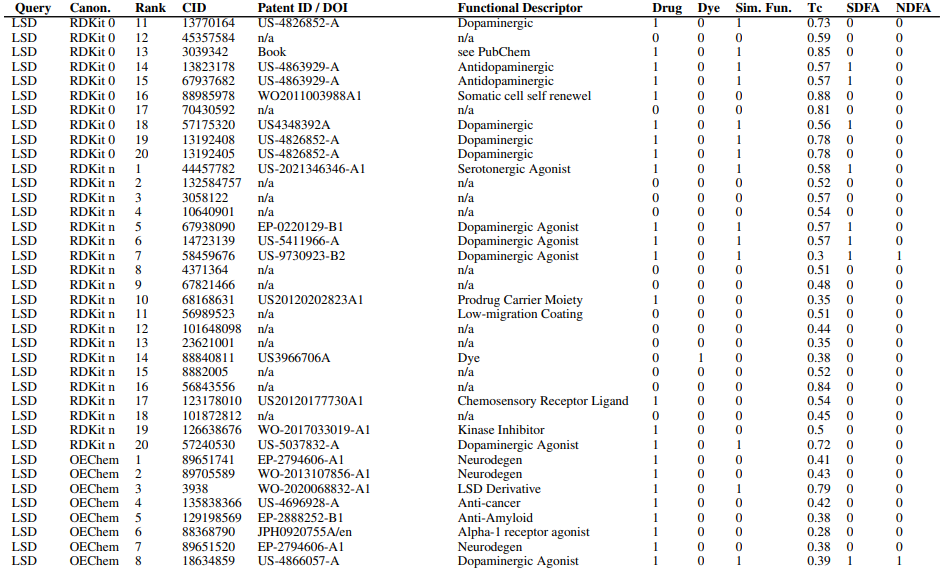
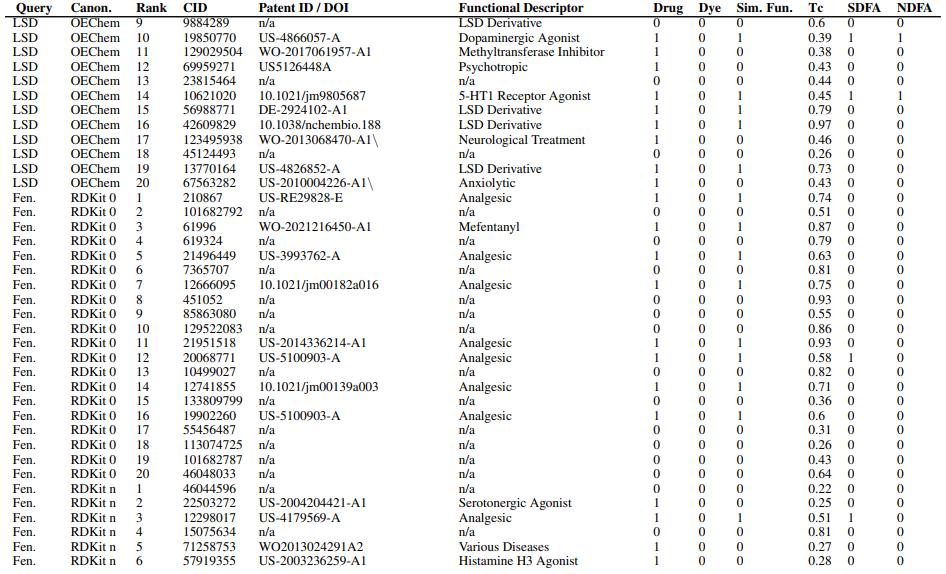
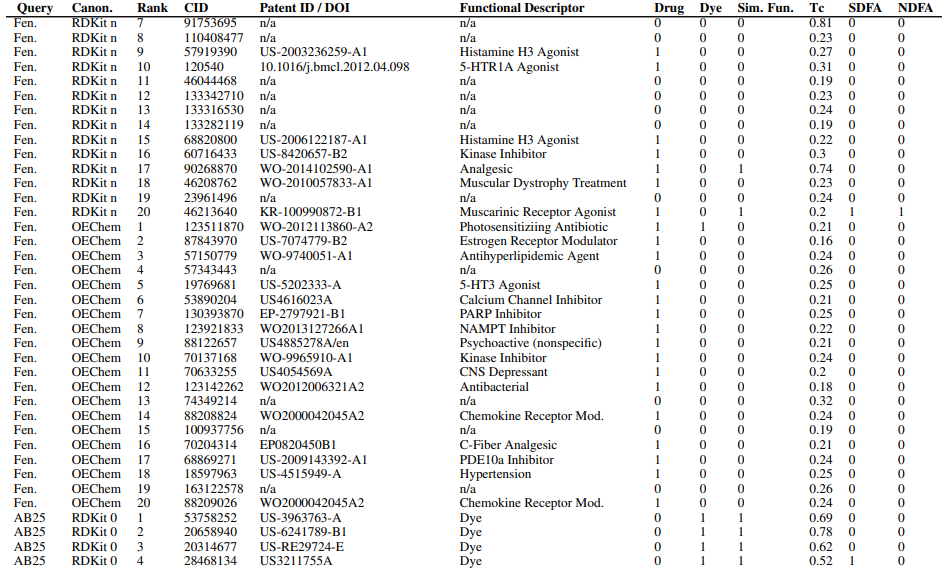
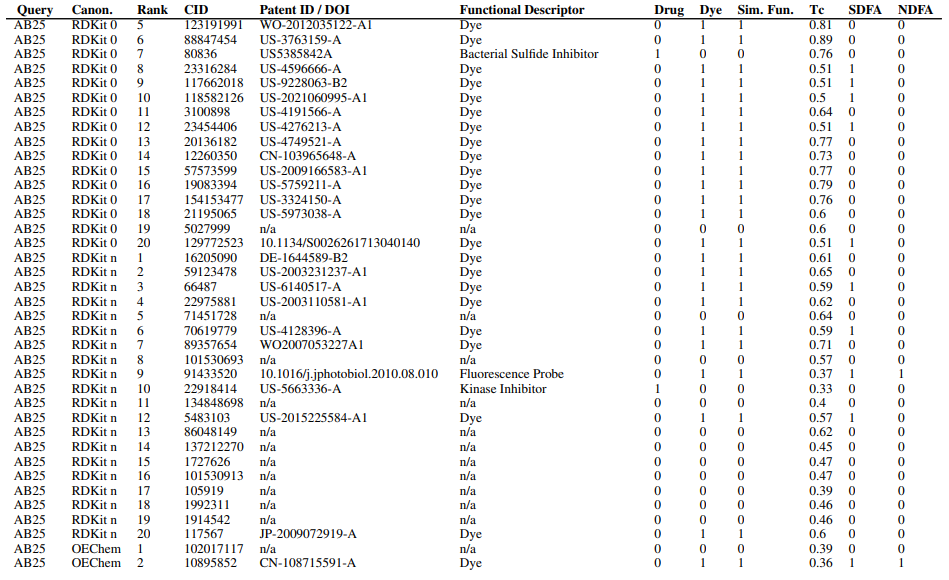
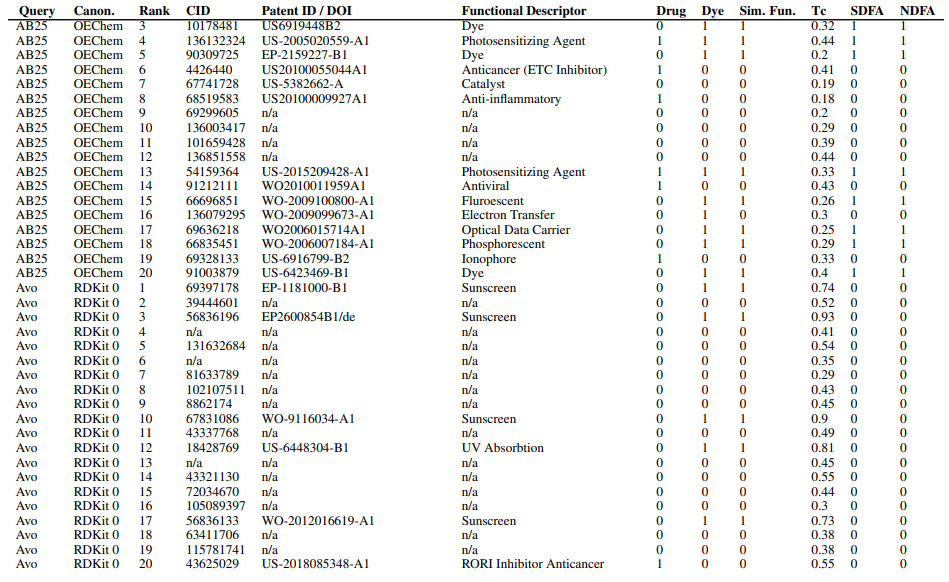
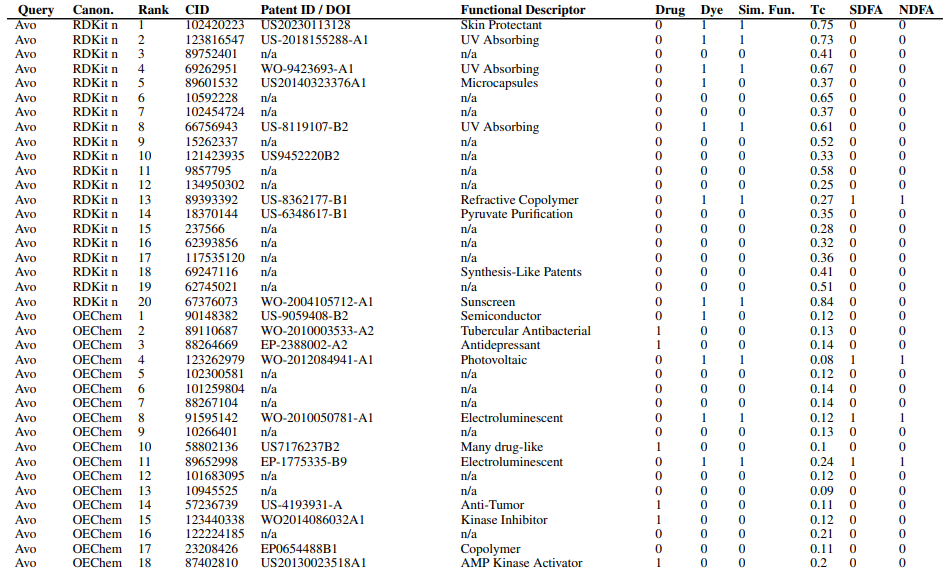
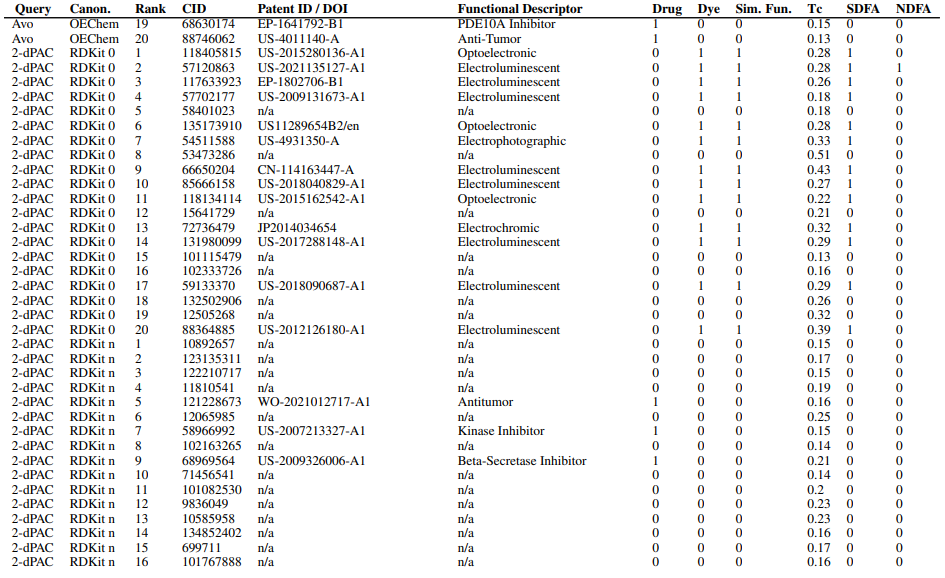
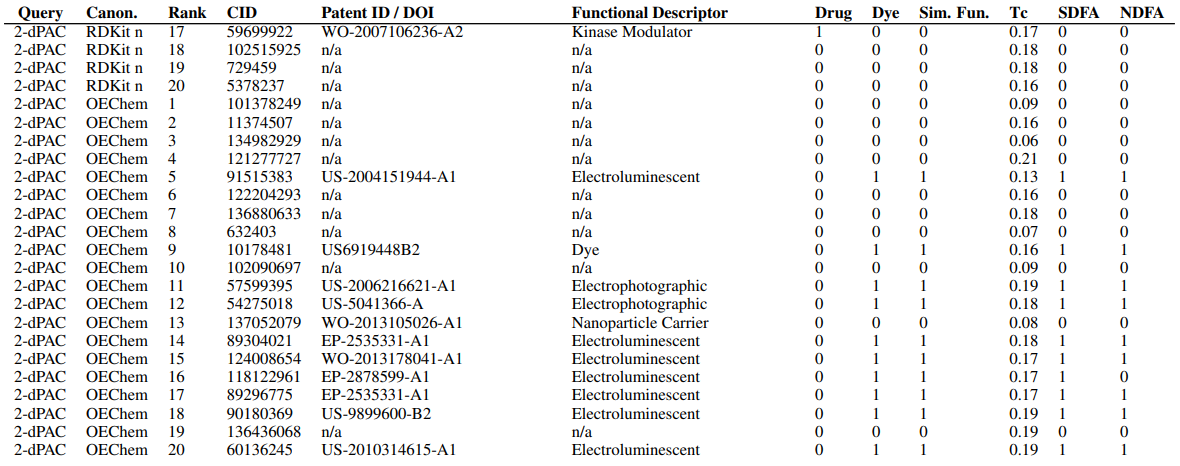
This paper is available on arxiv under CC BY-NC-SA 4.0 DEED license.
[story continues]
Written by
@penicillin
A tiny hero, Penicillin saves the day, infections yield to its might, transforming medicine's fight.
Topics and
tags
tags
ai-in-chemistry|chemical-similarity-search|drug-discovery|smiles|prompt-engineering|ai-research|molecular-embeddings|machine-learning
This story on HackerNoon has a decentralized backup on Sia.
Transaction ID: TSX_hmGFGGbLYANfiDHiLEHZV332B0LxkkTmczAs3R8
